Running from terminal#
There is an automatic script that will obtain focus from a folder containing a focus sequence.
If you have fits files you can simply run.
goodman-focus
It will run with the following defaults:
Argument |
Default Value |
Options |
|---|---|---|
|
Current Working Directory |
Any valid path |
|
*.fits |
Any |
|
gaussian |
moffat |
|
False |
True |
|
False |
True |
Where <input> is what you type.
To get some help and a full list of options use:
goodman-focus -h
Using it as a library#
After installing Install Using PYPI you can also import the class and instantiate it providing a set of arguments and values or using default ones.
from goodman_focus import GoodmanFocus
If no argument is provided it will instantiate with the default values.
The list of arguments can be defined as follow:
import os
from goodman_focus import GoodmanFocus
goodman_focus = GoodmanFocus(data_path=os.getcwd(),
file_pattern='*.fits',
obstype='FOCUS',
features_model='gaussian',
plot_results=False,
debug=False)
Which is equivalent to:
from goodman_focus import GoodmanFocus
goodman_focus = GoodmanFocus()
features_model is the function or model to fit to each detected line.
gaussian will use a Gaussian1D which provide more consistent results.
and moffat will use a Moffat1D model which fits the profile better but
is harder to control and results are less consistent than when using a gaussian.
Finally you need to call the instance, here is a full example.
from goodman_focus import GoodmanFocus
goodman_focus = GoodmanFocus()
results = goodman_focus()
However since version 0.3.0 you can pass a list of files and all will only check that all files exists
Interpreting Results#
The terminal version will print a message like this
Mode: IM__Red__VR Best Focus: -682.2891445722862 at FWHM: 2.610853168299203Best image: 0019_IM_FOCUS_VR-02-11-2019.fits with focus: -601 and FWHM: 2.618208314784383
Note
In version 2.0.0 21-10-2021 the format changed from the previous stable release in order to simplify the serialization process.
Using it as a library will return a list of dictionaries with the following values. Combination of settings for which the code is the same is called a mode, so the mode name is it’s unique identifier, how the name is constructed is explained in Decoding de mode name. In the following example only one object is included for simplicity.
[
{
"date": "2019-08-10",
"time": "2019-08-10T20:06:15.884",
"mode_name": "IM__Red__VR",
"focus": -837.3191595797898,
"fwhm": 2.5831939867345586,
"best_image_name": "0062_FOCUS_IMG_VR-10-08-2019.fits",
"best_image_focus": -799,
"best_image_fwhm": 2.6170559494587478,
"focus_data": [
-1994,
-1801,
-1601,
-1400,
-1201,
-997,
-799,
-601,
-400,
-200,
1
],
"fwhm_data": [
5.930451746635722,
5.306608292580763,
4.594963225848573,
3.555224927051857,
3.0090692421560217,
2.705695641392574,
2.6170559494587478,
2.7128327791088545,
3.02782407160001,
3.615781144676397,
4.265013680275817
]
}
]
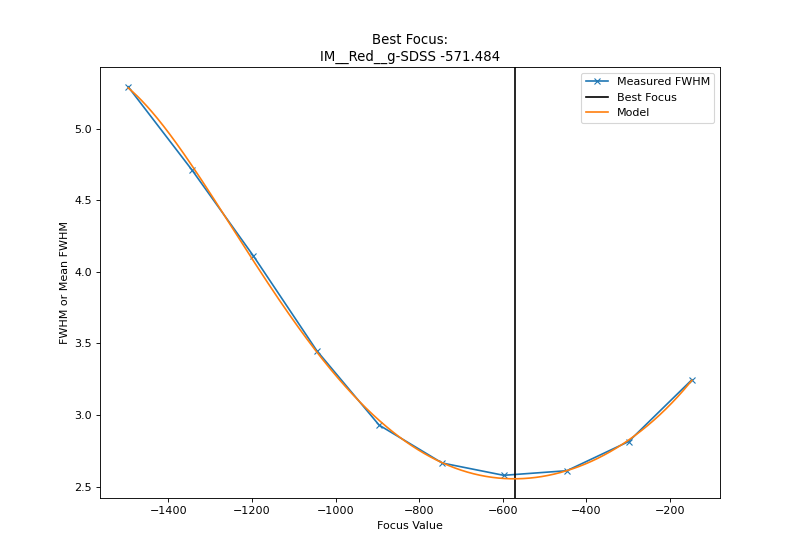
It is also possible to obtain a plot, from terminal, use --plot-results.
Below is a reproduction of results obtained with test data.
(Source code, png, hires.png, pdf)
Decoding de mode name#
The mode name is constructed using two letters to define the observing technique
(Imaging or Spectroscopy) and values obtained from the header. The characters
<, > and blanks are removed.
The mode name is different for Imaging and Spectroscopy, since for imaging the important settings are the instrument and the filter and for spectroscopy the important values come from the instrument, the grating and observing mode and filter from second filter wheel. Below, the word inside the parenthesis represents a keyword from the header.
Warning
Be aware that the separator string is a double underscore. This change
was necessary to avoid confusion with single underscores used in certain
keyword values.
For imaging:
IM__(INSTCONF)__(FILTER)
for example:
IM__Red__g-SDSS
For spectroscopy:
SP__(INSTCONF)__(WAVMODE)__(FILTER2)
for example:
SP__Red__400m2__GG455